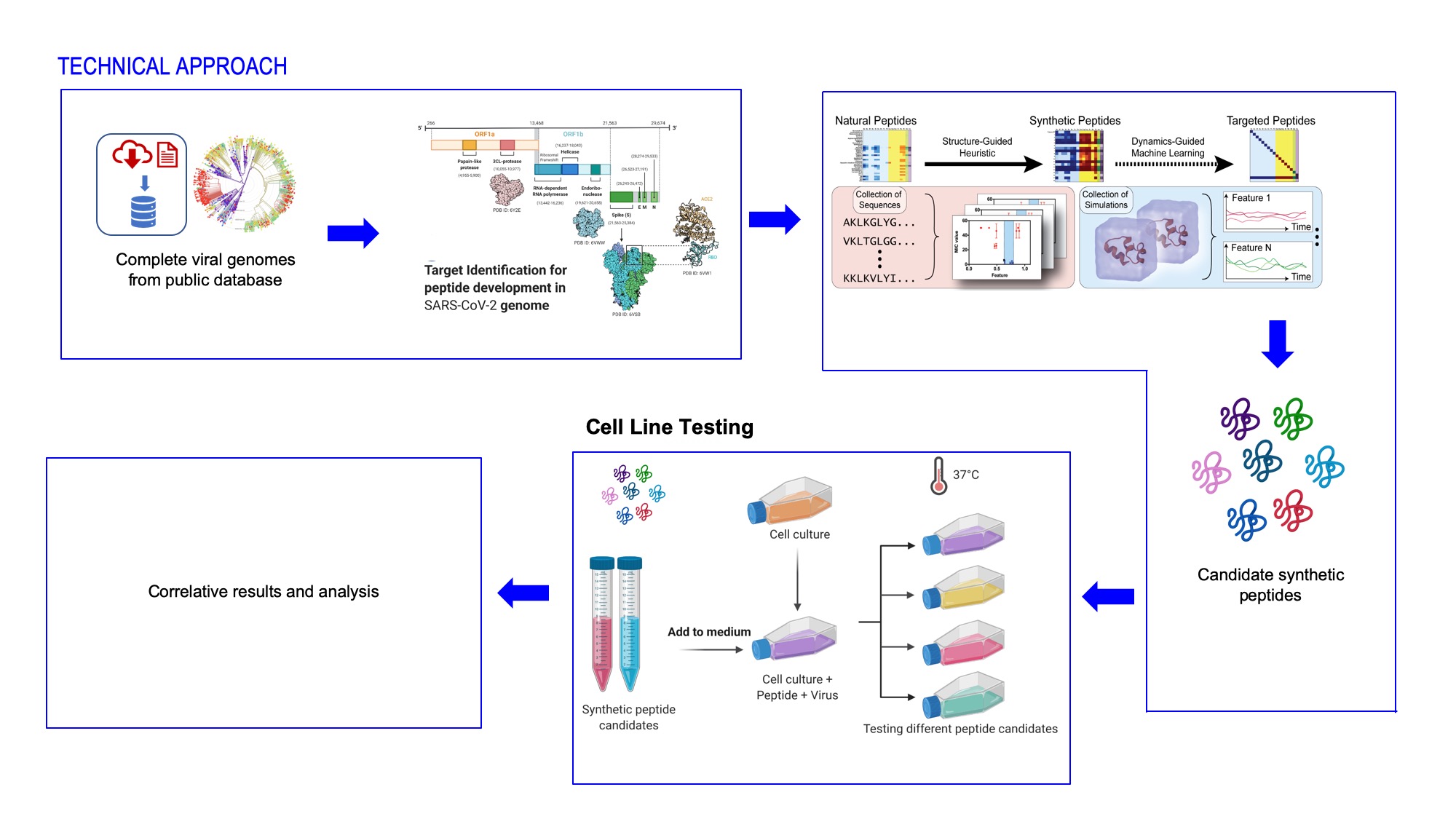
In this Viral Inactivation project, JPL is curating SARS nCoV-2 viral genomes to characterize various proteins, e.g., spike glycoproteins, polyproteins, and other viral replication proteins. At the direction of JPL, this COVID activity is performing computational analysis with Caltech collaborators. JPL-Caltech is using all the currently available SARS nCoV-2 genomes from the database and running it through JPL’s in-house SPROUT pipeline to predict 19 replication and virulent pathways. JPL in silico predicted antiviral peptide candidates will be produced at the Univ. Pennsylvania and then validated at the Univ. of Vermont Medical Center using live SARS nCoV-2 virus. This effort is ongoing with the goal of submitting the findings for peer review and publication.
VIRUS INACTIVATION - BROCHURE

-
Disclaimer
DISCLAIMER: The techniques and applications described herein have not been reviewed, cleared, or approved by FDA or other regulatory authority, nor have they received Coronavirus Disease 2019 (COVID-19) Emergency Use Authorizations for Medical Devices. Neither California Institute of Technology (including the Jet Propulsion Laboratory)("Caltech") nor its employees or agents provide any representation or warranty, express or implied, for fitness for a particular purpose, safety, efficacy, effectiveness, or non-infringement of any third party intellectual property rights. Caltech offers these application insights in good faith for others to build upon to help prevent the spread of COVID-19.
